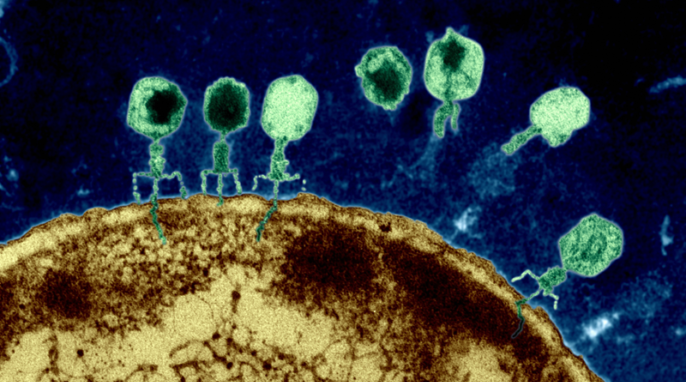
Malaria kills 1 million people each year, most of whom are children in sub-Saharan Africa. Turning to nature for new malaria drugs.
By Bill Sullivan, PhD
We live on a lush planet filled with over 290,000 species of plants. Herbs are a particular type of plant that lack a wooden stem, and humans have often sampled them in hopes of finding a new food or flavoring. Sometimes ingestion of an herb produces unwanted effects, such as death. But other herbs have medicinal qualities, such as the alleviation of fever.
An Old and New Malaria Drug
Dichroa febrifuga is one of the most important herbs in traditional Chinese medicine, used for millennia to treat ailments such as malaria. Malaria is caused by a unicellular parasite called Plasmodium that is transmitted by mosquitoes, and a high fever is one of the trademark symptoms.
Malaria continues to be a devastating disease, killing up to 1 million people each year, most of whom are children under the age of five in sub-Saharan Africa. There is an urgent need for new malaria drug treatments, since the parasite has developed resistance to most of our malaria drugs.
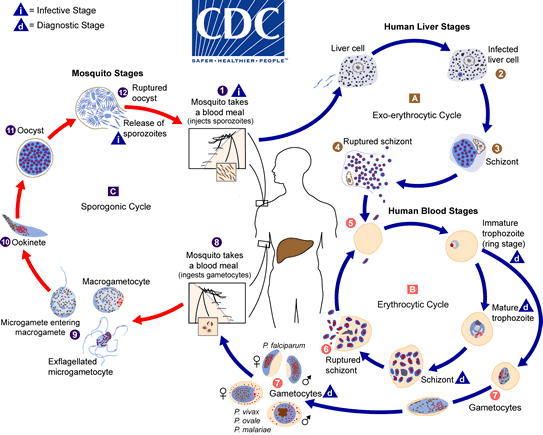
While effective against malaria, febrifugine is not tolerated well and has unwanted side effects. A better understanding of how febrifugine works is needed: How does it kill the malaria parasite? If the natural product’s mechanism of action against malaria could be identified, it would pave the way for the development of refined derivatives that are more specific against the parasite and less detrimental to patients. This has not been an easy task, but now scientists have identified an enzyme in the parasite that is inhibited by febrifugine. That enzyme is called prolyl-tRNA synthetase.
Prolyl-tRNA synthetase is critical for the production of proteins in a cell, a process known as translation. As shown in the figure, messenger RNA (mRNA), which serves as the “middleman” conveying the information in genes to build proteins, is read by molecular machines called ribosomes. Another type of RNA molecule called transfer RNA (tRNA) recognizes specific nucleotide sequences in the mRNA, bringing the corresponding amino acid to the ribosome so it can be added to a growing protein sequence.
RELATED: Sweet Solution: Taste Protein Guides Drug Design
Aminoacyl-tRNA synthetase enzymes are needed to “charge” the tRNA; in other words, they attach the correct amino acid to the correct tRNA. When prolyl-tRNA synthetase is blocked by febrifugine, the amino acid proline does not get attached to tRNA. This leads to a buildup of “uncharged” tRNA, which is interpreted as a sign of starvation by the cell—or in this case by the single-celled malaria parasite. Proline is a common amino acid needed to build many proteins, and when prolyl-tRNA synthetase cannot do its job, protein production grinds to a halt. Even better, this enzyme is required in multiple stages of the parasite’s life cycle, so both the liver and the blood forms of the parasite are impacted.
But as mentioned earlier, humans do not tolerate febrifugine very well, probably because we also have a version of prolyl-tRNA synthetase and perhaps other proteins that febrifugine poisons. Identification of this drug target is helping researchers develop derivatives of febrifugine, such as halofuginol, that act more strongly against the parasite’s prolyl-tRNA synthetase with less toxicity in humans.
So how did scientists figure out that febrifugine targets prolyl-tRNA synthetase? There are several ways to identify the molecular mechanism of drug activity. In this case, the researchers cultured malaria in the presence of the drug, forcing the parasites to evolve or die. Those that lived were less sensitive to febrifugine, meaning that they accrued a genetic change (one or more mutations in their DNA) that allowed them to persist despite the presence of the drug. This process is analogous to the development of penicillin-resistant bacteria.
RELATED: Making Antipsychotics with Baker’s Yeast
Scientists sequenced the genomes of the parasites that were able to grow better in febrifugine. Such a feat would have taken many years and millions of dollars not long ago, but today it has become routine. The genome sequence of the febrifugine-resistant parasites contained a common mutation in the gene encoding prolyl-tRNA synthetase, which signaled that this enzyme plays a critical role in the drug’s action. Understanding how the parasite develops resistance also helps scientists design compounds that act on the target differently. As you may surmise, we are in a constant “arms race” with these insidious microbes, but this discovery is a step toward a victory for us.
About the Author
Dr. Bill Sullivan is a professor of Pharmacology and Toxicology, and of Microbiology and Immunology, at the Indiana University School of Medicine. He studies the biology of parasites as a means to develop new treatments for infectious disease. Dr. Sullivan routinely blogs about science and pop culture at The ‘Scope. Follow Dr. Sullivan on X, formerly Twitter, @wjsullivan.
Reference
Herman, J. D.; Pepper, L. R.; Cortese, J. F.; Estiu, G.; Galinsky, K.; Zuzarte-Luis, V.; Derbyshire, E. R.; Ribacke, U.; Lukens, A. K.; Santos, S. A.; Patel, V.; Clish, C. B.; Sullivan, W. J. Jr.; Zhou, H.; Bopp, S. E.; Schimmel, P.; Lindquist, S.; Clardy, J.; Mota, M. M.; Keller, T. L.; Whitman, M.; Wiest, O.; Wirth, D. F.; Mazitschek, R. (2015). The cytoplasmic prolyl-tRNA synthetase of the malaria parasite is a dual-stage target of febrifugine and its analogs. Science Translational Medicine, 7 (288) PMID: 25995223.
A version of this article appeared previously on The ‘Scope.

![Dichroa febrifuga, a medicinal herb that has been historically used to treat fever, is named for its active ingredient, febrifugine. By Keith Edkins (Own work) [CC BY-SA 3.0], via Wikimedia Commons](https://magazine.scienceconnected.org/wp-content/uploads/2016/05/Dichroa_febrifuga_Glasgow-e1463012377212.jpg)